Introduction to R Packages, Markdown and Notebooks
Last updated on 2026-04-23 | Edit this page
Overview
Questions
- What is an
R package? - How to install
Rpackages? - What is
R MarkdownandR Notebooks? - How can I integrate my
Rcode with text and plots? - How can I convert
.Rmdfiles to.html?
Objectives
- Understand what an
R packageis - Install packages using the
packagestab. - Install packages using
Rcode. - Understand basic syntax of
R MarkdownandR Notebooks
Acknowledgement
This workshop was adapted using material from the Data Carpentry
lessons R for Social Scientists,
specifically lesson 00-intro
and lesson 06-rmarkdown.
Other Materials
What are R packages?
R Packages are the
fundamental units of reproducible R code. They are
collections of reusable R functions, sample data, and the
documentation that describes how to use the functions.
What is the difference between base R and
packages?
The base R package
contains the basic functions which let R function as a
language:
- Arithmetic
- Input/output
- Basic programming support, etc
The R software is distributed with the
base R package installed. In addition to the
base R installation, there are in excess of 20,000
additional packages which can be used to extend the functionality of
R. Many of these have been written by R users
and have been made available in central repositories, like the one
hosted at the Comprehensive R Archive Network CRAN,
for anyone to download and install into their own R
environment.
CRAN
is a network of ftp and web servers around the world that store
identical, up-to-date, versions of code and documentation for
R.
Installing packages using R code and the
packages tab
We’ll use the tidyverse and here packages
in this workshop.
You can install these packages from the console by typing the command
install.packages(), or from the packages
tab.
We’ll install tidyverse from the console, and
here from the packages tab.
R
install.packages("tidyverse")
OUTPUT
The following package(s) will be installed:
- tidyverse [2.0.0]
These packages will be installed into "/__w/irim-r-workshops/irim-r-workshops/renv/profiles/lesson-requirements/renv/library/linux-ubuntu-noble/R-4.5/x86_64-pc-linux-gnu".
# Installing packages --------------------------------------------------------
[32m✔[0m tidyverse 2.0.0 [linked from cache]
Successfully installed 1 package in 3.6 milliseconds.You can see if you have a package installed by looking in the
packages tab (on the lower-right by default). You can also
type the command installed.packages() into the console and
examine the output.
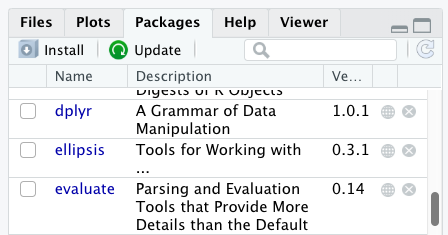
Packages can also be installed from the packages tab. On
the packages tab, click the Install icon and
start typing the name of the package you want in the text box. As you
type, packages matching your starting characters will be displayed in a
drop-down list so that you can select them.
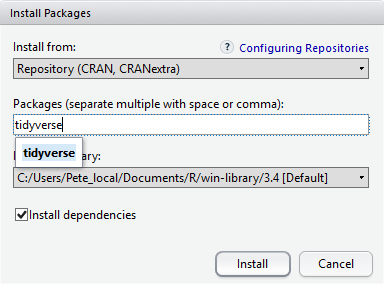
At the bottom of the Install Packages window is a check
box to Install dependencies. This is ticked by default,
which is usually what you want. Packages can (and do) make use of
functionality built into other packages, so for the functionality
contained in the package you are installing to work properly, there may
be other packages which have to be installed with them. The
Install dependencies option makes sure that this
happens.
Exercise
Use the Packages tab to confirm that you have both the
tidyverse and here packages installed.
Scroll through packages tab down to tidyverse. You can
also type a few characters into the searchbox. The
tidyverse package is really a package of packages,
including ggplot2 and dplyr, both of which
require other packages to run correctly. All of these packages will be
installed automatically. Depending on what packages have previously been
installed in your R environment, the install of tidyverse
could be very quick or could take several minutes. As the install
proceeds, messages relating to its progress will be written to the
console. You will be able to see all of the packages which are actually
being installed.
Because the install process accesses the CRAN
repository, you will need an Internet connection to install
packages.
It is also possible to install packages from other repositories, as
well as Github or the local file system, but we won’t be
looking at these options in this workshop.
R Markdown and R Notebooks
R Markdown is a flexible type of document that allows
you to seamlessly combine executable R code, and its
output, with text in a single document.
An R Notebook is a specific interactive execution mode
for an R Markdown (Rmd) document. Code chunks
are executed independently and interactively within the
RStudio editor.
R Markdown documents can be readily converted to
multiple static and dynamic output formats, including PDF (.pdf), Word
(.docx), and HTML (.html).
The benefit of a well-prepared R Markdown or
Notebook document is full reproducibility. This also means
that, if you notice a data transcription error, or you are able to add
more data to your analysis, you will be able to recompile the report
without making any changes in the actual document.
Creating an R Notebook file
To create a new R Markdown document in
RStudio, click
File -> New File -> R Notebook. You may be prompted
to install required packages the first time you do this.
Basic components of an R Notebook
YAML Header
To control the output, a YAML (YAML Ain’t
Markup Language) header is needed:
---
title: "My Awesome Report"
output: html_document
---The header is defined by the three hyphens at the beginning
(---) and the three hyphens at the end
(---).
In the YAML, the only required field is the
output:, which specifies the type of output you want. This
can be an html_document, a pdf_document, or a
word_document. We will start with an HTML document and
discuss the other options later.
After the header, to begin the body of the document, you start typing
after the end of the YAML header (i.e. after the second
---).
Markdown syntax
Markdown is a popular markup language that allows you to
add formatting elements to text, such as bold,
italics, and code. The formatting will not be
immediately visible in a markdown (.md) document, like you would see in
a Word document. Rather, you add Markdown syntax to the
text, which can then be converted to various other files that can
translate the Markdown syntax. Markdown is
useful because it is lightweight, flexible, and platform
independent.
RStudio provides a real time preview of the formatting-
click the Visual tab to view the rendered
Markdown, or Source to view the raw
Markdown.
Headings
A # in front of text indicates to Markdown
that this text is a heading. Adding more #s make the
heading smaller, i.e. one # is a first level heading, two
##s is a second level heading, etc. up to the 6th level
heading.
# Title
## Section
### Sub-section
#### Sub-sub section
##### Sub-sub-sub section
###### Sub-sub-sub-sub section(only use a level if the one above is also in use)
Formatting
You can make things bold by surrounding the word
with double asterisks, **bold**, or double underscores,
__bold__; and italicize using single asterisks,
*italics*, or single underscores,
_italics_.
You can also combine bold and italics to
write something really important with
triple-asterisks, ***really***, or underscores,
___really___; and, if you’re feeling bold (pun intended),
you can also use a combination of asterisks and underscores,
**_really_**, **_really_**.
To create code-type font, surround the word with
backticks, `code-type`.
Code Chunks
Code chunks are blocks where you write and execute R code. They start
with ```{r} and end with ```.
To insert a Chunk, click the small arrow next to the
Insert button in the editor toolbar and select
R.
To run a Chunk, click the small green play arrow on the right side of
the chunk, or use the keyboard shortcut
Ctrl+Alt+I on Windows and
Linux (or Cmd+Option+I on
Mac).
Render and Share Your Notebook
Once your analysis is complete, you can generate a final, polished report.
Click the Preview (or Render) button in the
RStudio editor toolbar.
This creates a self-contained HTML file (or PDF/Word document,
depending on your settings in your YAML header) that
includes both the narrative text and the final results.
You can easily share this output file with others, even if they don’t
use R.
Now that we’ve learned a couple of things, it might be useful to implement them.
Create your own new R Notebook
Start by opening a new R Notebook:
Click File -> New File -> R Notebook
When you open a new R Notebook, some explanatory text is
provided. This can be deleted so you can enter your own text and
code.
Download data
We will be using a dataset called SAFI_clean.csv. The
direct download link for this file is: https://github.com/datacarpentry/r-socialsci/blob/main/episodes/data/SAFI_clean.csv.
This data is a slightly cleaned up version of the
SAFI Survey Results available on figshare.
First, we need to create a new folder called data to
store this dataset. Go to the Files pane, and create a new folder named
data, and two subfolders called cleaned and
raw.
intro_r
│
└── scripts
│
└── data
│ └── cleaned
│ └── raw
│
└─── images
│
└─── documentsYou can either download the SAFI_clean.csv dataset used
for this workshop from the GitHub link or with R. You can
download the file from this GitHub link
and save it as SAFI_clean.csv in the data/raw
directory you just created. Or you can do this directly from
R by copying and pasting this in your console:
download.file( "https://raw.githubusercontent.com/datacarpentry/r-socialsci/main/episodes/data/SAFI_clean.csv", "data/raw/SAFI_clean.csv", mode = "wb" )
Start an Introduction section
Make a header called Introduction, and insert some
explanatory text about the dataset that will be in your report. For
example:
This report uses the tidyverse package
along with the SAFI dataset, which has columns
that include:
- village
- interview_date
- no_members
- years_liv
- respondent_wall_type
- roomsYou can also create an ordered list using numbers:
1. village
2. interview_date
3. no_members
4. years_liv
5. respondent_wall_type
6. roomsAnd nested items by tab-indenting:
- village
- Name of village
- interview_date
- Date of interview
- no_members
- How many family members lived in a house
- years_liv
- How many years respondent has lived in village or neighbouring
village
- respondent_wall_type
- Type of wall of house
- rooms
- Number of rooms in houseFor more Markdown syntax see the following reference guide.
Now we can render the document into HTML by clicking the
preview button in the top of the Source
pane (top left). If you haven’t saved the document yet, you will be
prompted to do so when you preview for the
first time.
Writing an R Markdown report
Now we will add some R code to demonstrate (we will
learn more about this code in the next workshop!).
First, we need to make sure tidyverse
is loaded. It is not enough to load
tidyverse from the console, we will need
to load it within our R Notebook. The same applies to our
data. To load these, we will need to create a ‘code chunk’ at the top of
our document (below the YAML header).
A code chunk can be inserted by clicking
Code \> Insert Chunk, or by using the keyboard shortcuts
Ctrl+Alt+I on Windows and
Linux, and Cmd+Option+I on
Mac.
The syntax of a code chunk is:
An R Markdown document knows that this text is not part
of the report from the (```) that begins and ends the
chunk. It also knows that the code inside of the chunk is R code from
the r inside of the curly braces ({}). After
the r you can add a name for the code chunk . Naming a
chunk is optional, but recommended. Each chunk name must be unique, and
only contain alphanumeric characters and -.
To load tidyverse and our
SAFI_clean.csv file, we will insert a chunk and call it
‘setup’. Since we don’t want this code or the output to show in our
rendered HTML document, we add an include = FALSE option
after the code chunk name ({r setup, include = FALSE}).
MARKDOWN
```{r setup, include = FALSE}
library(tidyverse)
library(here)
interviews <- read_csv(here("data/raw/SAFI_clean.csv"), na = "NULL")
```Important Note!
The file paths you give in a .Rmd document, e.g. to load a .csv file, are relative to the .Rmd document, not the project root.
We highly recommend the use of the here() function to
keep the file paths consistent within your project.
Insert table
Next, we will create a table which shows the average household size
grouped by village and memb_assoc. We can do
this by creating a new code chunk and calling it ‘interview-tbl’. Or,
you can come up with something more creative (just remember to stick to
the naming rules).
We will learn more about this code later!
To see the output, run the code chunk with the green triangle in the
top right corner of the the chunk, or with the keyboard shortcuts:
Ctrl+Alt+C on Windows and
Linux, or Cmd+Option+C on
Mac.
To make sure the table is formatted nicely in our output document, we
will need to use the kable() function from the
knitr package. The kable()
function takes the output of your R code and knits it into a nice
looking HTML table. You can also specify different aspects of the table,
e.g. the column names, a caption, etc.
Run the code chunk to make sure you get the desired output.
R
interviews %>%
filter(!is.na(memb_assoc)) %>%
group_by(village, memb_assoc) %>%
summarize(mean_no_membrs = mean(no_membrs)) %>%
knitr::kable(caption = "We can also add a caption.",
col.names = c("Village", "Member Association",
"Mean Number of Members"))
| Village | Member Association | Mean Number of Members |
|---|---|---|
| Chirodzo | no | 8.062500 |
| Chirodzo | yes | 7.818182 |
| God | no | 7.133333 |
| God | yes | 8.000000 |
| Ruaca | no | 7.178571 |
| Ruaca | yes | 9.500000 |
Many different R packages can be used to generate
tables. Some of the more commonly used options are listed in the table
below.
| Name | Creator(s) | Description |
|---|---|---|
| condformat | Oller Moreno (2022) | Apply and visualize conditional formatting to data frames in R. It renders a data frame with cells formatted according to criteria defined by rules, using a tidy evaluation syntax. |
| DT | Xie et al. (2023) | Data objects in R can be rendered as HTML tables using the JavaScript library ‘DataTables’ (typically via R Markdown or Shiny). The ‘DataTables’ library has been included in this R package. |
| formattable | Ren and Russell (2021) | Provides functions to create formattable vectors and data frames. ‘Formattable’ vectors are printed with text formatting, and formattable data frames are printed with multiple types of formatting in HTML to improve the readability of data presented in tabular form rendered on web pages. |
| flextable | Gohel and Skintzos (2023) | Use a grammar for creating and customizing pretty tables. The following formats are supported: ‘HTML’, ‘PDF’, ‘RTF’, ‘Microsoft Word’, ‘Microsoft PowerPoint’ and R ‘Grid Graphics’. ‘R Markdown’, ‘Quarto’, and the package ‘officer’ can be used to produce the result files. |
| gt | Iannone et al. (2022) | Build display tables from tabular data with an easy-to-use set of functions. With its progressive approach, we can construct display tables with cohesive table parts. Table values can be formatted using any of the included formatting functions. |
| huxtable | Hugh-Jones (2022) | Creates styled tables for data presentation. Export to HTML, LaTeX, RTF, ‘Word’, ‘Excel’, and ‘PowerPoint’. Simple, modern interface to manipulate borders, size, position, captions, colours, text styles and number formatting. |
| pander | Daróczi and Tsegelskyi (2022) | Contains some functions catching all messages, ‘stdout’ and other useful information while evaluating R code and other helpers to return user specified text elements (e.g., header, paragraph, table, image, lists etc.) in ‘pandoc’ markdown or several types of R objects similarly automatically transformed to markdown format. |
| pixiedust | Nutter and Kretch (2021) | ‘pixiedust’ provides tidy data frames with a programming interface intended to be similar to ’ggplot2’s system of layers with fine-tuned control over each cell of the table. |
| reactable | Lin et al. (2023) | Interactive data tables for R, based on the ‘React Table’ JavaScript library. Provides an HTML widget that can be used in ‘R Markdown’ or ‘Quarto’ documents, ‘Shiny’ applications, or viewed from an R console. |
| rhandsontable | Owen et al. (2021) | An R interface to the ‘Handsontable’ JavaScript library, which is a minimalist Excel-like data grid editor. |
| stargazer | Hlavac (2022) | Produces LaTeX code, HTML/CSS code and ASCII text for well-formatted tables that hold regression analysis results from several models side-by-side, as well as summary statistics. |
| tables | Murdoch (2022) | Computes and displays complex tables of summary statistics. Output may be in LaTeX, HTML, plain text, or an R matrix for further processing. |
| tangram | Garbett et al. (2023) | Provides an extensible formula system to quickly and easily create production quality tables. The processing steps are a formula parser, statistical content generation from data defined by a formula, and rendering into a table. |
| xtable | Dahl et al. (2019) | Coerce data to LaTeX and HTML tables. |
| ztable | Moon (2021) | Makes zebra-striped tables (tables with alternating row colors) in LaTeX and HTML formats easily from a data.frame, matrix, lm, aov, anova, glm, coxph, nls, fitdistr, mytable and cbind.mytable objects. |
Customizing chunk output
We mentioned using include = FALSE in a code chunk to
prevent the code and output from printing in the knitted document. There
are additional options available to customize how the code-chunks are
presented in the output document. The options are entered in the code
chunk after chunk-name and separated by commas, e.g.
{r chunk-name, eval = FALSE, echo = TRUE}.
| Option | Options | Output |
|---|---|---|
eval |
TRUE or FALSE
|
Whether or not the code within the code chunk should be run. |
echo |
TRUE or FALSE
|
Choose if you want to show your code chunk in the output document.
echo = TRUE will show the code chunk. |
include |
TRUE or FALSE
|
Choose if the output of a code chunk should be included in the
document. FALSE means that your code will run, but will not
show up in the document. |
warning |
TRUE or FALSE
|
Whether or not you want your output document to display potential warning messages produced by your code. |
message |
TRUE or FALSE
|
Whether or not you want your output document to display potential messages produced by your code. |
fig.align |
default, left, right,
center
|
Where the figure from your R code chunk should be output on the page |
Exercise
Play around with the different options in the chunk with the code for the table, and see what each option does to the output.
What happens if you use eval = FALSE and
echo = FALSE? What is the difference between this and
include = FALSE?
Create a chunk with {r eval = FALSE, echo = FALSE}, then
create another chunk with {r include = FALSE} to compare.
eval = FALSE and echo = FALSE will neither run
the code in the chunk, nor show the code in the knitted document. The
code chunk essentially doesn’t exist in the rendered document as it was
never run. Whereas include = FALSE will run the code and
store the output for later use.
In-line R code
Now we will use some in-line R code to present some
descriptive statistics. To use in-line R code, we use the
same backticks that we used in the Markdown section, with
an r to specify that we are generating R-code. The
difference between in-line code and a code chunk is the number of
backticks. In-line R code uses one backtick
(`r`), whereas code chunks use three backticks
(```r```).
For example, today’s date is `r Sys.Date()`, will be
rendered as: today’s date is 2026-04-23.
The code will display today’s date in the output document (well,
technically the date the document was last knitted or previewed).
The best way to use in-line R code, is to minimize the amount of code you need to produce the in-line output by preparing the output in code chunks. Let’s say we’re interested in presenting the average household size in a village.
R
# create a summary data frame with the mean household size by village
mean_household <- interviews %>%
group_by(village) %>%
summarize(mean_no_membrs = mean(no_membrs))
# and select the village we want to use
mean_chirodzo <- mean_household %>%
filter(village == "Chirodzo")
Now we can make an informative statement on the means of each village, and include the mean values as in-line R-code. For example:
The average household size in the village of Chirodzo is
`r round(mean_chirodzo$mean_no_membrs, 2)`
becomes…
The average household size in the village of Chirodzo is 7.08.
Because we are using in-line R code instead of the
actual values, we have created a dynamic document that will
automatically update if we make changes to the dataset and/or code
chunks.
Plots
Finally, we will also include a plot, so our document is a little more colourful and a little less boring. We will create some code to use in the plotting.
R
interviews_plotting <- interviews %>%
## pivot wider by items_owned
separate_rows(items_owned, sep = ";") %>%
## if there were no items listed, changing NA to no_listed_items
replace_na(list(items_owned = "no_listed_items")) %>%
mutate(items_owned_logical = TRUE) %>%
pivot_wider(names_from = items_owned,
values_from = items_owned_logical,
values_fill = list(items_owned_logical = FALSE)) %>%
## pivot wider by months_lack_food
separate_rows(months_lack_food, sep = ";") %>%
mutate(months_lack_food_logical = TRUE) %>%
pivot_wider(names_from = months_lack_food,
values_from = months_lack_food_logical,
values_fill = list(months_lack_food_logical = FALSE)) %>%
## add some summary columns
mutate(number_months_lack_food = rowSums(select(., Jan:May))) %>%
mutate(number_items = rowSums(select(., bicycle:car)))
R
interviews_plotting %>%
ggplot(aes(x = respondent_wall_type)) +
geom_bar(aes(fill = village))
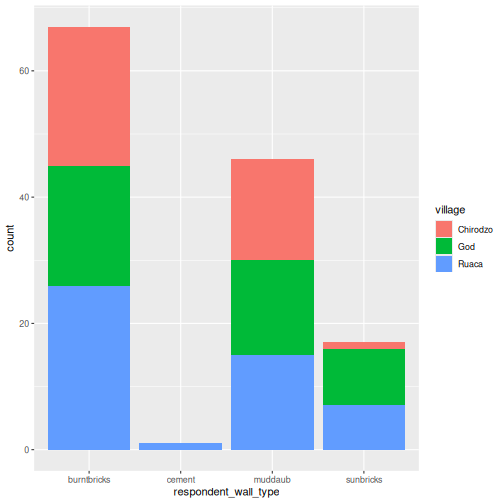
We can also create a caption with the chunk option
fig.cap.
R
interviews_plotting %>%
ggplot(aes(x = respondent_wall_type)) +
geom_bar(aes(fill = village), position = "dodge") +
labs(x = "Type of Wall in Home", y = "Count", fill = "Village Name") +
scale_fill_viridis_d() # add colour deficient friendly palette
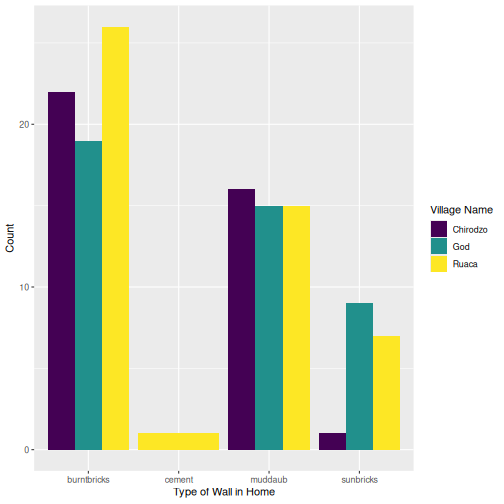
Other output options
You can convert R Markdown to a PDF or a Word document
(among others). Click the little triangle next to the
Preview button to get a drop-down menu. Or
you could put pdf_document or word_document in
the initial header of the file.
---
title: "My Awesome Report"
author: "Author name"
date: ""
output: word_document
---Note: Creating PDF documents
Creating .pdf documents may require installation of some
extra software. The R package tinytex provides some tools
to help make this process easier for R users. With tinytex
installed, run tinytex::install_tinytex() to install the
required software (you’ll only need to do this once) and then when you
Knit to pdf tinytex will
automatically detect and install any additional LaTeX packages that are
needed to produce the pdf document. Visit the tinytex website for
more information.
Note: Inserting citations into an
R Markdown file
It is possible to insert citations into an R Markdown
file using the editor toolbar. The editor toolbar includes commonly seen
formatting buttons generally seen in text editors (e.g., bold and italic
buttons). The toolbar is accessible by using the settings dropdown menu
(next to the Preview dropdown menu) to select
Use Visual Editor, also accessible through the shortcut
Crtl+Shift+F4. From here, clicking
Insert allows Citation to be selected
(shortcut: Crtl+Shift+F8). For example,
searching 10.1007/978-3-319-24277-4 in
From DOI and inserting will provide the citation for
ggplot2 [@wickham2016]. This will also save
the citation(s) in ‘references.bib’ in the current working directory.
Visit the R Studio website
for more information. Tip: obtaining citation information from relevant
packages can be done by using citation("package").
Resources
R MarkdowndocumentationR Markdown cheat sheetGetting started with R MarkdownIntroduction to R Markdown-
R Markdown: The Definitive Guide(book byRstudioteam)
- Use
install.packages()to install packages (libraries) - Use
library()to load packages -
R Markdownis a useful language for creating reproducible documents combining text and executableRcode - Specify chunk options to control formatting of the output document
